library(pipeline)
dir <- pipeline_skeleton(dir = tempfile(), single = TRUE)Warning
This package:
- is a work-in-progress, proof-of-concept;
- is unstable in both it’s API and output;
- may not fully fledge past the prototype stage.
Background
The pipeline package’s aim is to makes it easy for users to adopt a functional approach to their analysis workflows. Pipelines are defined using a reduced subset of R and, when run, use an intelligent approach to work planning and object caching. It was motivated by a desire to reduce the boilerplate I was writing with my R scripts and associated Makefiles. It can be thought of a simpler, but less capable, alternative to targets. Whilst the goal is to be “good enough” for many pieces of analysis, by encouraging a functional approach, it remains easy to move to more advanced alternatives.
Scope
Goals:
- 🗸 Intelligent caching to reduce unnecessary execution.
- 🗸 Configuration files using a reduced subset of base R as their syntax.
- 🗸 Simple and efficient approach to managing multiple scenarios.
Out of scope:
- 🗴 HPC orchestration.
- 🗴 Automatic parallelisation of the pipeline.
A motivating example
The easiest way to get started with the package is to look at some of the included examples. pipeline_skeleton() can be used to set up a pipeline based on the mtcars data set and across single or multiple scenarios. We start with the single scenario example which has been carefully constructed to illustrate the different functionality that pipeline has.
#> ✔ Template pipeline created in '/tmp/Rtmp7aN1tO/file539d1d2fbb5a'.We now have the following in dir:
#> /tmp/Rtmp7aN1tO/file539d1d2fbb5a
#> ├── R
#> │ ├── load_dat.R
#> │ ├── plot_dat.R
#> │ └── wrangle_dat.R
#> ├── config.R
#> ├── data
#> │ └── mtcars.csv
#> └── pipeline.RThe pipeline we are going to run is described in pipeline.R:
pipeline.R
out <- pipeline_run({
raw <- load_dat(CONFIG$in_csv)
clean <- wrangle_dat(raw, CONFIG$rows)
plot <- plot_dat(clean, CONFIG$out_plot)
})pipeline_run() takes an expression where each assignment it contains represents a stage of the analysis. There are a few things to note about the expression syntax:
- The left-hand-side (target) of each assignment must be uniquely named.
- Function calls on the right-hand-side (rhs) must be defined in the function directory
R, contained in the base R namespace or, if present in another package, be explicitly namespaced (i.e.package::fun). - The target names cannot match the names of functions in base R, or those contained in the function directory.
- The order of assignments is not important as the order of orchestration is determined by
pipeline_run(). - Values from the configuration file
config.Rare accessed by the specialCONFIGvalue.
The configuration file
Configuration files are simply R scripts that are evaluated in a dedicated environment with some allowances for multiple scenarios that we discuss at the end.
Within this environment we enable two additional special functions INFILE and OUTFILE:
INFILE()marks a file as an input. This ensures thatpipeline_run()knows to keep a hash of the object and to monitor it for future changes when the pipeline is rerun.OUTFILE()marks a file name as an output. This ensures that outputs from different scenarios (see later) do not overwrite one another (output files get saved in a scenario-dependent directory). It also ensures that output files are recreated if they are deleted.
In our example we have the following configuration file:
config.r
in_csv <- INFILE("data/mtcars.csv")
out_plot <- OUTFILE("plot.png")
rows <- 10Here in_csv points to our input data and out_plot to a png output file.
The R directory
This is the directory where users define functions that represent stages of their analysis (e.g. cleaning data, modelling, plotting, …). Here we have:
R/load_dat.R
load_dat <- function(file) {
# Note: If function not in the base namespace or function directory
# we must explictly namespace the call using `::`.
utils::read.csv(file)
}This file contains a single function which takes the filename of raw data as input before doing a little wrangling and returning the output. Note the comment about namespacing functions; anything not in the base R namespace or the function directory needs to be explicitly namespaced.
Next we have R/wrangle_dat.R:
R/wrangle_dat.R
wrangle_dat <- function(dat, rows) {
stopifnot(is.data.frame(dat), nrow(dat) >= rows, rows >= 0)
head(dat, rows)
}This function curtails the number of rows of data. Here we also make use of assertions in the function, checking that dat is a data frame and rows is between 0 and the number of rows in the data frame (inclusive). Assertions like these can be thought of unit tests for your analysis. Whilst here we have them at the start of the function to check inputs, they can be used anywhere in your function to ensure things are running as expected. If an assertion fails it will stop the pipeline running, bringing your attention to any issue at hand.
Finally there is
R/plot_dat.R
This function takes the manipulated data as input and creates a plot which is saved as a png. When saving output to disk, we recommend that the function returns the location where the file is saved for later use.
Putting it together
Running the pipeline we see the following
original_directory <- setwd(dir)
out <- pipeline_run({
raw <- load_dat(CONFIG$in_csv)
clean <- wrangle_dat(raw, CONFIG$rows)
plot <- plot_dat(clean, CONFIG$out_plot)
})#>
#> ── Starting pipeline with `default` configuration ──────────────────────────────
#> ℹ Running function to generate object `raw`.
#> ℹ Saving `raw` to disk.
#> ℹ Running function to generate object `clean`.
#> ℹ Saving `clean` to disk.
#> ℹ Running function to generate object `plot`.
#> ℹ Saving `plot` to disk.Each target output is saved within a single list that we have allocated to out. The name of the list is configurable but, by default, is ‘default’.
str(out$default$raw)
#> 'data.frame': 32 obs. of 12 variables:
#> $ car : chr "Mazda RX4" "Mazda RX4 Wag" "Datsun 710" "Hornet 4 Drive" ...
#> $ mpg : num 21 21 22.8 21.4 18.7 18.1 14.3 24.4 22.8 19.2 ...
#> $ cyl : int 6 6 4 6 8 6 8 4 4 6 ...
#> $ disp: num 160 160 108 258 360 ...
#> $ hp : int 110 110 93 110 175 105 245 62 95 123 ...
#> $ drat: num 3.9 3.9 3.85 3.08 3.15 2.76 3.21 3.69 3.92 3.92 ...
#> $ wt : num 2.62 2.88 2.32 3.21 3.44 ...
#> $ qsec: num 16.5 17 18.6 19.4 17 ...
#> $ vs : int 0 0 1 1 0 1 0 1 1 1 ...
#> $ am : int 1 1 1 0 0 0 0 0 0 0 ...
#> $ gear: int 4 4 4 3 3 3 3 4 4 4 ...
#> $ carb: int 4 4 1 1 2 1 4 2 2 4 ...
str(out$default$clean)
#> 'data.frame': 10 obs. of 12 variables:
#> $ car : chr "Mazda RX4" "Mazda RX4 Wag" "Datsun 710" "Hornet 4 Drive" ...
#> $ mpg : num 21 21 22.8 21.4 18.7 18.1 14.3 24.4 22.8 19.2
#> $ cyl : int 6 6 4 6 8 6 8 4 4 6
#> $ disp: num 160 160 108 258 360 ...
#> $ hp : int 110 110 93 110 175 105 245 62 95 123
#> $ drat: num 3.9 3.9 3.85 3.08 3.15 2.76 3.21 3.69 3.92 3.92
#> $ wt : num 2.62 2.88 2.32 3.21 3.44 ...
#> $ qsec: num 16.5 17 18.6 19.4 17 ...
#> $ vs : int 0 0 1 1 0 1 0 1 1 1
#> $ am : int 1 1 1 0 0 0 0 0 0 0
#> $ gear: int 4 4 4 3 3 3 3 4 4 4
#> $ carb: int 4 4 1 1 2 1 4 2 2 4
out$default$plot
#> [1] "/tmp/Rtmp7aN1tO/file539d1d2fbb5a/output/default/plot.png"
#> attr(,"OUTFILE")
#> [1] TRUE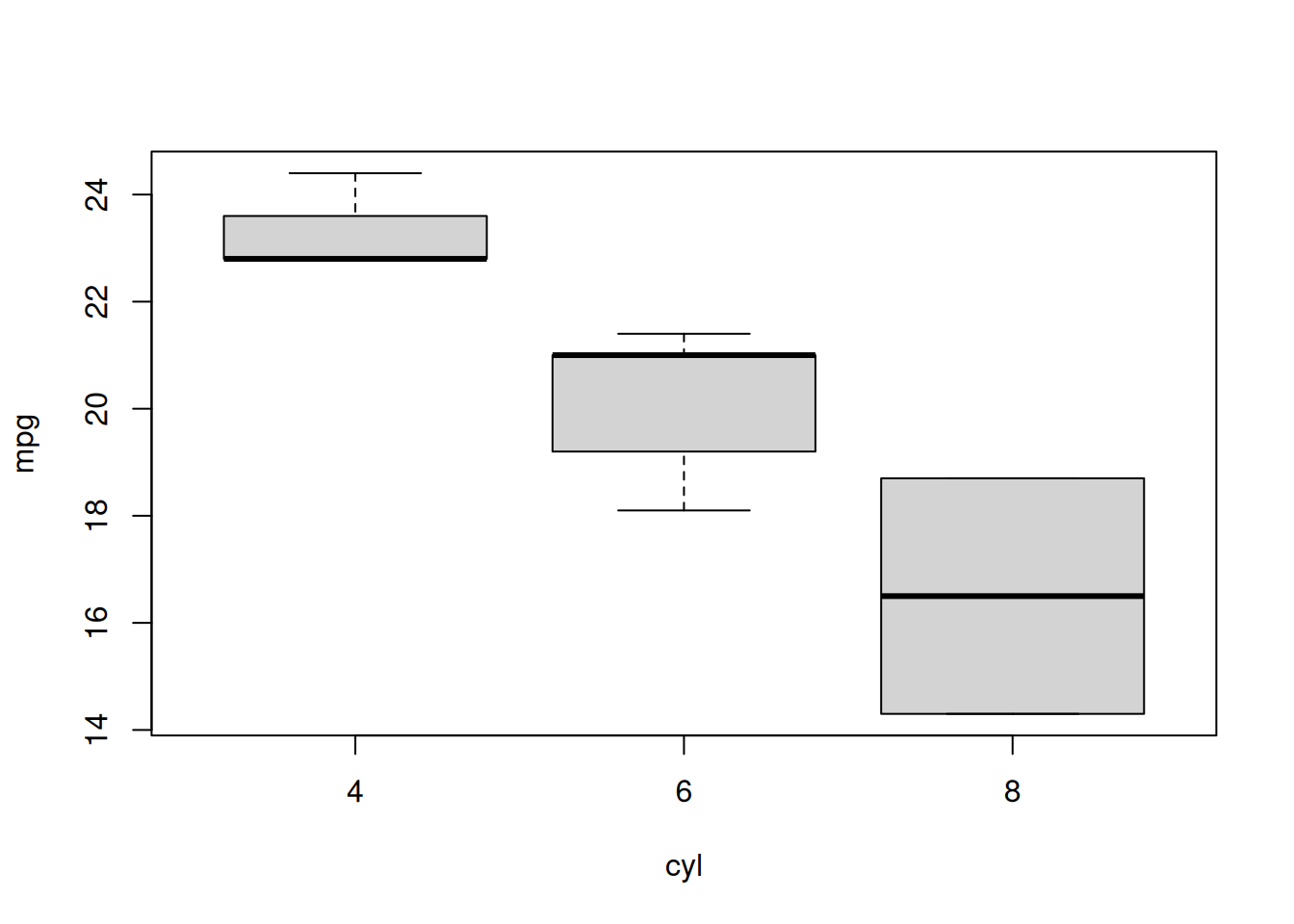
The outputs, along with some additional information that we can use for cache invalidation checks, are also saved to disk in the folder ./pipeline_cache. We do not expect uses to interact directly with this folder and it’s contents should not be altered.
#> /tmp/Rtmp7aN1tO/file539d1d2fbb5a/.pipeline_cache
#> ├── 6d95a1c536b565f927cc7c5e9edfb17a.rds
#> ├── 94433eb0504ad33a41c79b71e3eb2920.rds
#> ├── bc84fd5137211c57b156a5dfbc4e1914.rds
#> └── default_cache_.rdsChanging things
Let’s assume that after cleaning our data we wanted to sample rows rather than take as is. Updating our pipeline accordingly, we have
out <- pipeline_run({
raw <- load_dat(CONFIG$in_csv)
clean <- wrangle_dat(raw, CONFIG$rows)
samples <- clean[sample(nrow(clean), replace = TRUE), ]
plot <- plot_dat(samples, CONFIG$out_plot)
})#>
#> ── Starting pipeline with `default` configuration ──────────────────────────────
#> ℹ Loading object `raw` from disk cache.
#> ℹ Loading object `clean` from disk cache.
#> ℹ Running function to generate object `samples`.
#> ℹ Saving `samples` to disk.
#> ℹ Running function to generate object `plot`.
#> ℹ Saving `plot` to disk.Note that where possible, rather than start from scratch, the cached results were loaded from disk. This caching takes account of changes in functions within the function directory but does not attempt to handle changes in the underlying R version or packages you use (in essence, it assumes environment stability is managed elsewhere).
Multiple scenarios
The package assists in running multiple scenarios in two ways. Firstly, users can specify different configurations to run in the call, i.e.
pipeline_run(
...,
config = c("default", "production"),
scenario_default = "default"
)This in turn tells the function to parse the configuration file, looking for an entry called ‘default’. This is used as the base scenario. Other values (in this case ‘production’) in the configuration file are then layered on top of this (internally using modifyList()) to generate additional scenarios to run.
Secondly, we want to avoid unnecessary computations so, where possible, we try to share cached objects across scenarios. To make things more concrete, consider our second demo:
dir2 <- pipeline_skeleton(dir = tempfile(), single = FALSE)
setwd(dir2)#> ✔ Template pipeline created in '/tmp/Rtmp7aN1tO/file539d581f0a49'.This time the configuration file defines two scenarios, default, which subsets the data to 10 rows and production, which returns 32 rows. Note that when working across different scenarios, the entries in the configuration file must be appropriately named lists.
config.R
To run both scenarios:
out <- pipeline_run(
{
raw <- load_dat(CONFIG$in_csv)
clean <- wrangle_dat(raw, CONFIG$rows)
plot <- plot_dat(clean, CONFIG$out_plot)
},
scenario = c("default", "production"),
scenario_default = "default" # included here but note that this is
# also the default value for the argument.
)#>
#> ── Starting pipeline with `default` configuration ──────────────────────────────
#> ℹ Running function to generate object `raw`.
#> ℹ Saving `raw` to disk.
#> ℹ Running function to generate object `clean`.
#> ℹ Saving `clean` to disk.
#> ℹ Running function to generate object `plot`.
#> ℹ Saving `plot` to disk.
#>
#> ── Starting pipeline with `production` configuration ───────────────────────────
#> ℹ Loading object `raw` from disk cache.
#> ℹ Running function to generate object `clean`.
#> ℹ Saving `clean` to disk.
#> ℹ Running function to generate object `plot`.
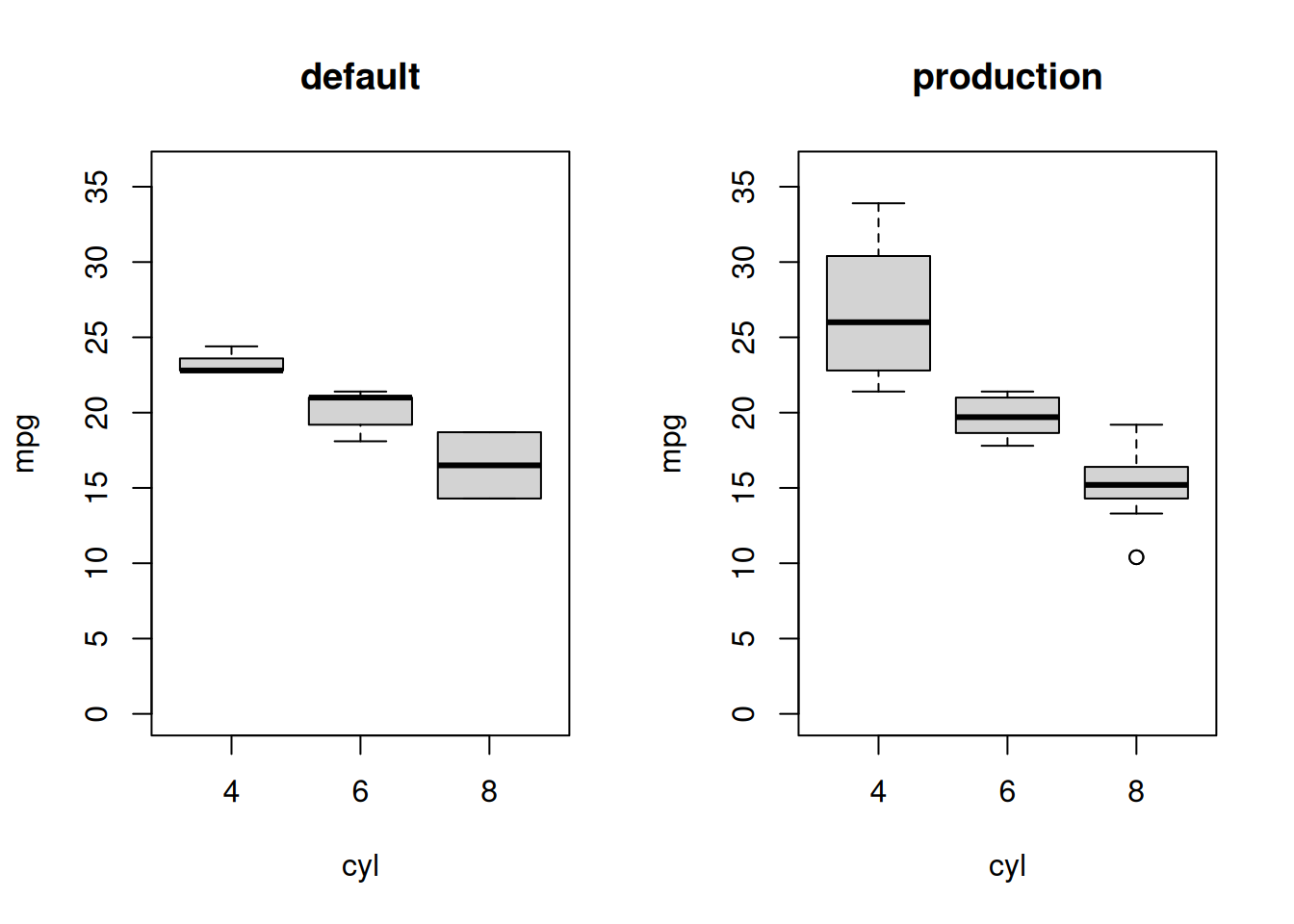
#> ℹ Saving `plot` to disk.We see above that the default scenario ran from scratch where as the production scenario was able to skip the first computation, loading the cached result from disk.